PDF) Multiple Trait Covariance Association Test Identifies Gene Ontology Categories Associated with Chill Coma Recovery Time in Drosophila melanogaster

Proteome-wide association studies identify biochemical modules associated with a wing-size phenotype in Drosophila melanogaster | Nature Communications
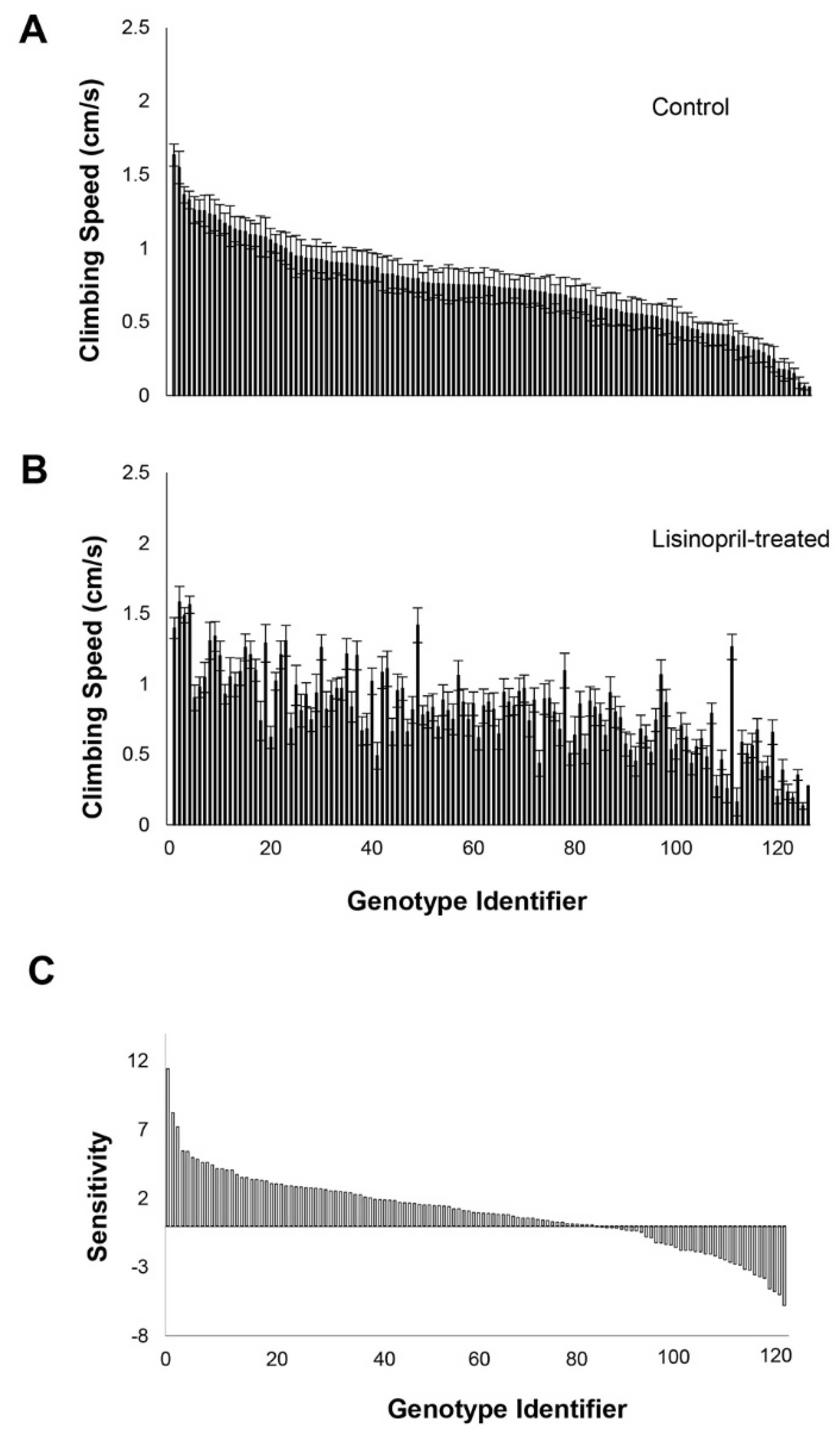
Genes | Free Full-Text | Genome-Wide Analysis in Drosophila Reveals the Genetic Basis of Variation in Age-Specific Physical Performance and Response to ACE Inhibition

PDF) Nucleotide diversity inflation as a genome-wide response to experimental lifespan extension in Drosophila melanogaster

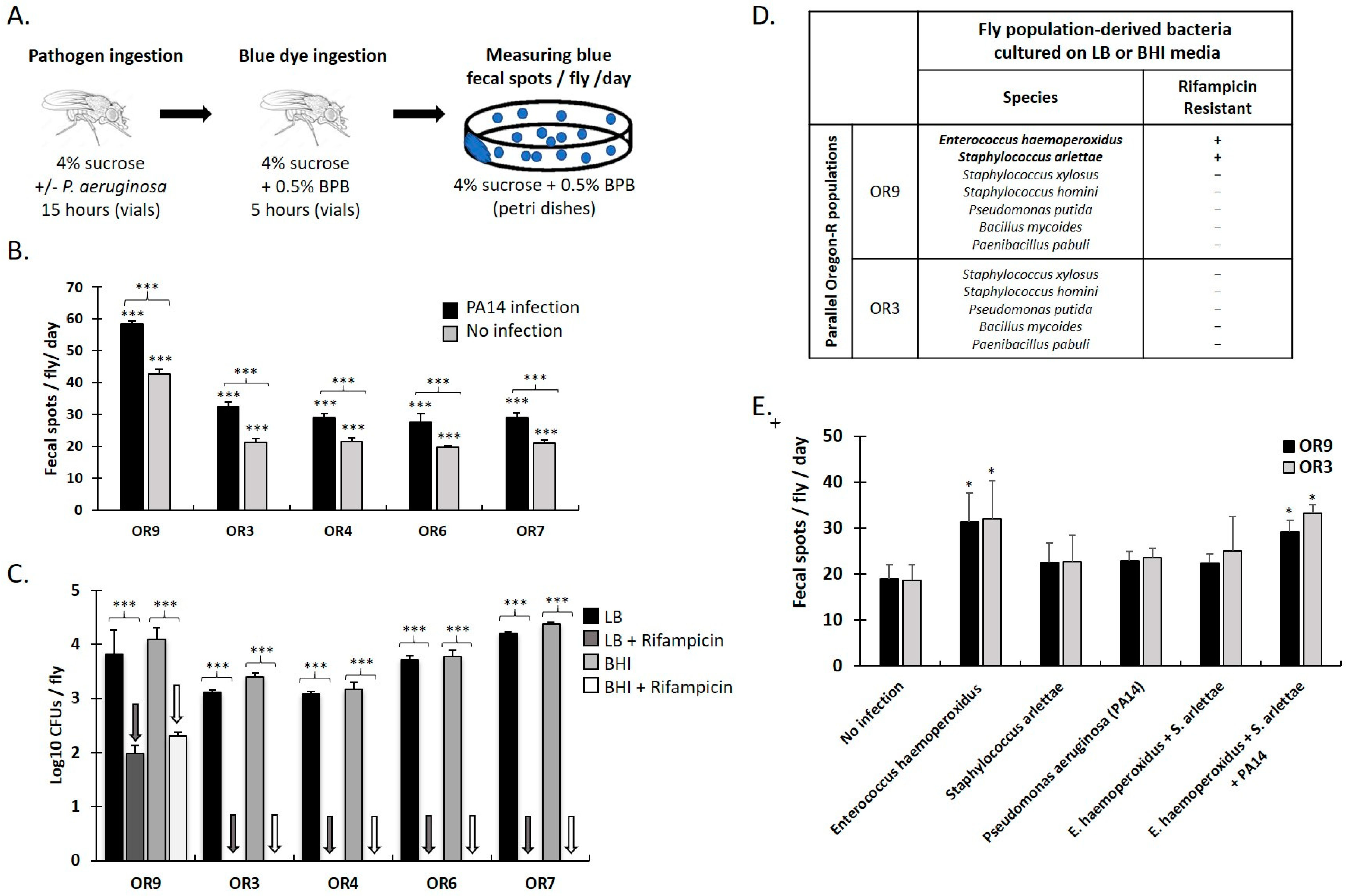
Commensal bacteria act as a broad genetic buffer in Drosophila during chronic under-nutrition | bioRxiv
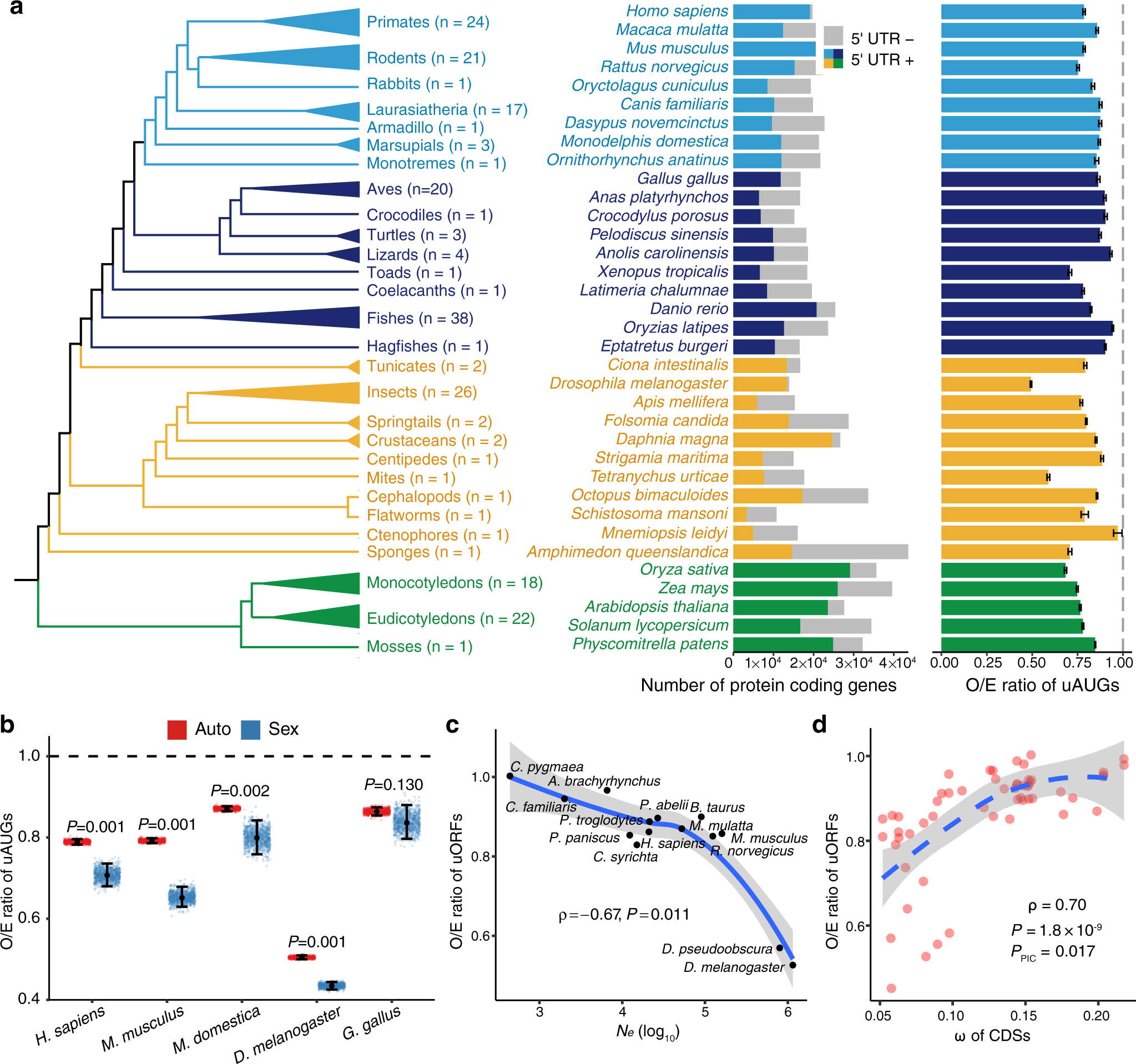
Determinants of genome-wide distribution and evolution of uORFs in eukaryotes | Nature Communications

Nacα protects the larval fat body from cell death by maintaining cellular proteostasis in Drosophila | Nature Communications

A generalizable deep learning framework for inferring fine-scale germline mutation rate maps | Nature Machine Intelligence

PDF) Genome Wide Association Studies of early fitness traits in Drosophila melanogaster unveil plasticity and decoupling of different aspects of phenotype
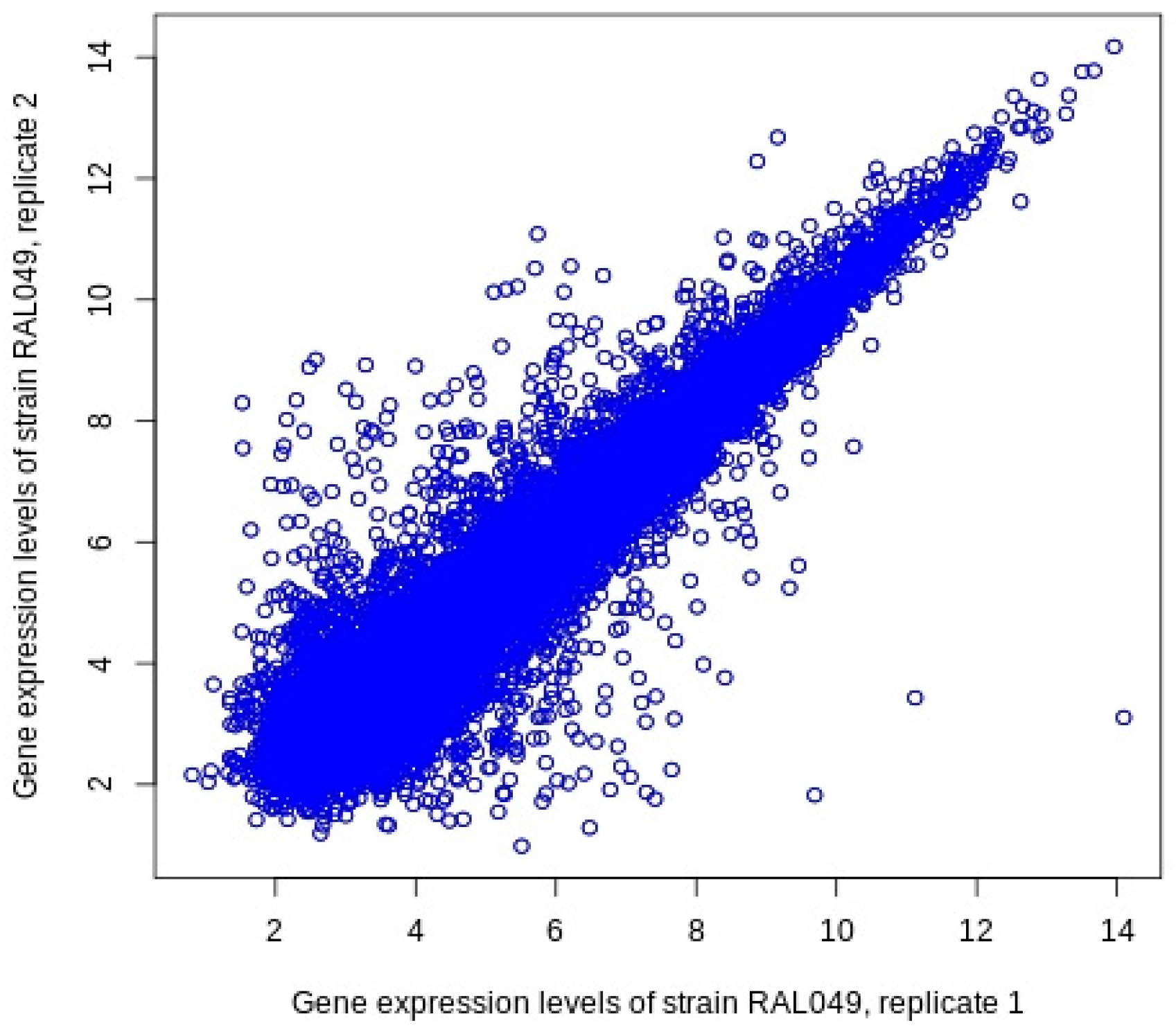
BioMedInformatics | Free Full-Text | Identifying Genes Related to Retinitis Pigmentosa in Drosophila melanogaster Using Eye Size and Gene Expression Data

Natural variation in the regulation of neurodevelopmental genes modifies flight performance in Drosophila | bioRxiv
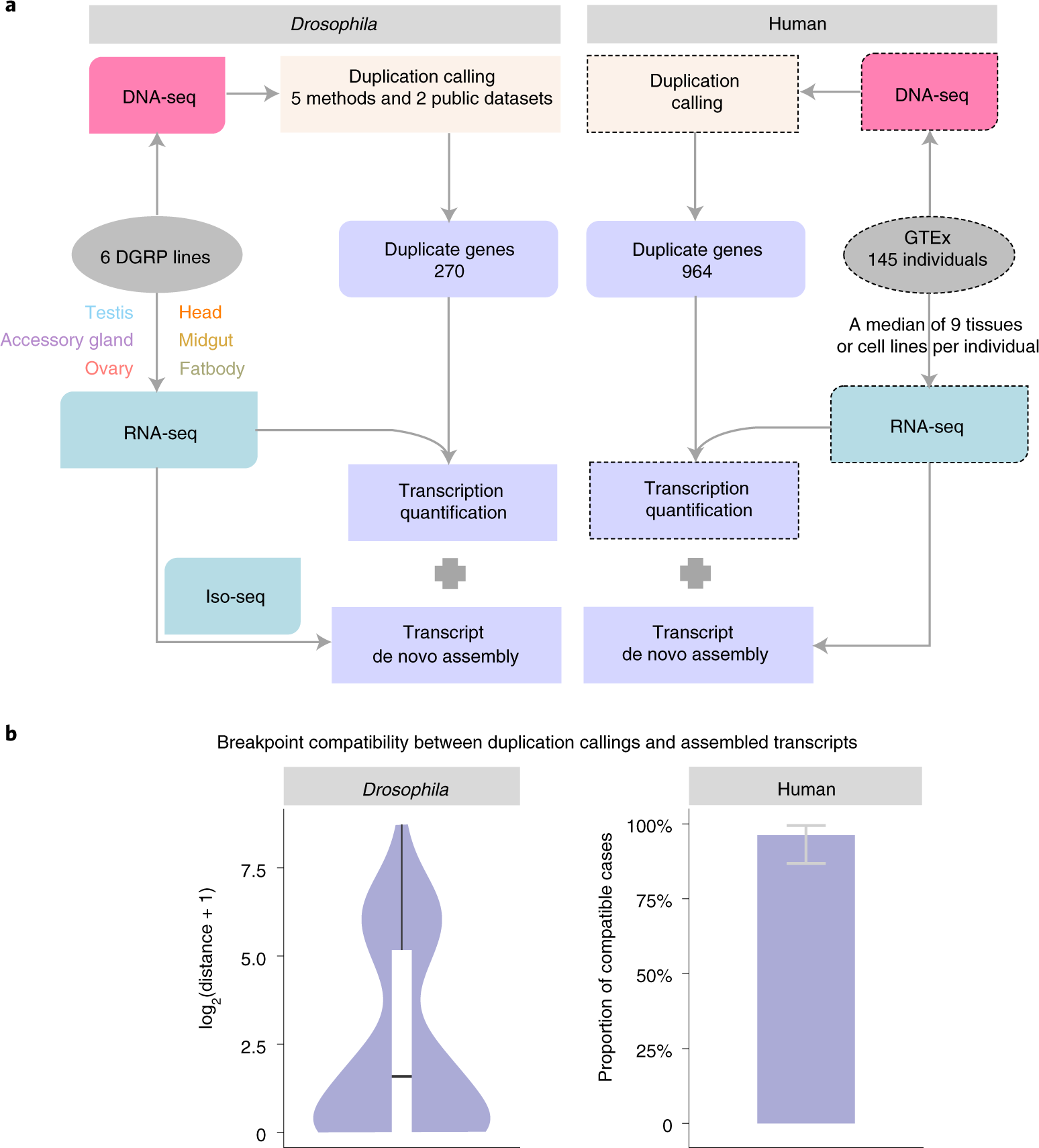
Dosage sensitivity and exon shuffling shape the landscape of polymorphic duplicates in Drosophila and humans | Nature Ecology & Evolution

Transcriptional changes in specific subsets of Drosophila neurons following inhibition of the serotonin transporter | Translational Psychiatry

Sex‐dependent and sex‐independent regulatory systems of size variation in natural populations | Molecular Systems Biology
Natural variation in the regulation of neurodevelopmental genes modifies flight performance in Drosophila | PLOS Genetics